|
Zhao Yang (杨钊) I am a fourth-year PhD student at the Gaoling School of Artificial Intelligence, Renmin University of China, supervised by Prof. Bing Su. I also work closely with Prof. Chuan Cao. My research interests focus on Language Models and Diffusion Models, with a particular emphasis on their applications in AI4Science. I expect to graduate in 2027 and will be entering the job market. Feel free to reach out! |

|
Research
|
Preprints |
|
|
D3LM
|
D3LM: A Discrete DNA Diffusion Language Model for Bidirectional DNA Understanding and Generation
Zhao Yang*, Hengchang Liu*, Chuan Cao, Bing Su arXiv / MLGenX Workshop, 2026 Paper / model We present D3LM, a discrete DNA diffusion language model that unifies bidirectional DNA understanding and DNA generation. |
|
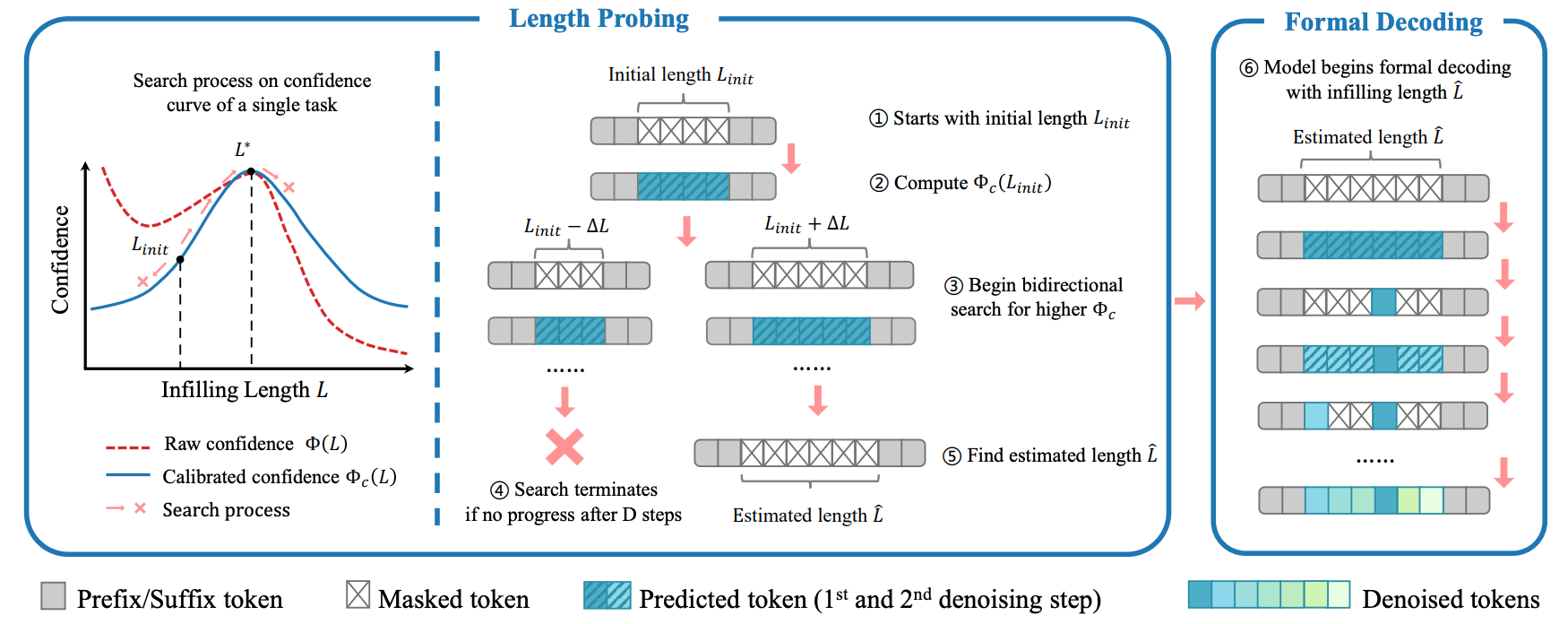
Diffusion LMs Can Approximate Optimal Infilling Lengths Implicitly
Hengchang Liu*, Zhao Yang*, Bing Su arXiv, 2026 Paper / code We investigate the ability of diffusion-based language models to implicitly determine the optimal infilling lengths. |
|
NatureLM
|
Nature Language Model: Deciphering the Language of Nature for Scientific Discovery
Yingce Xia, Peiran Jin, Shufang Xie, Liang He, Chuan Cao, ..., Zhao Yang, ..., Tao Qin arXiv, 2025 Paper / project A sequence-based science foundation model that unifies molecules, proteins, DNA and RNA for cross-domain scientific discovery. |
|
HybriDNA
|
HybriDNA: A Hybrid Transformer-Mamba2 Long-Range DNA Language Model
Mingqian Ma, Guoqing Liu, Chuan Cao, Pan Deng, Tri Dao, Albert Gu, Peiran Jin, Zhao Yang, Yingce Xia, Renqian Luo, Pipi Hu, Zun Wang, Yuan-Jyue Chen, Haiguang Liu, Tao Qin arXiv, 2025 Paper / project A hybrid Transformer-Mamba2 long-context DNA language model for genomic understanding and generation. |
|
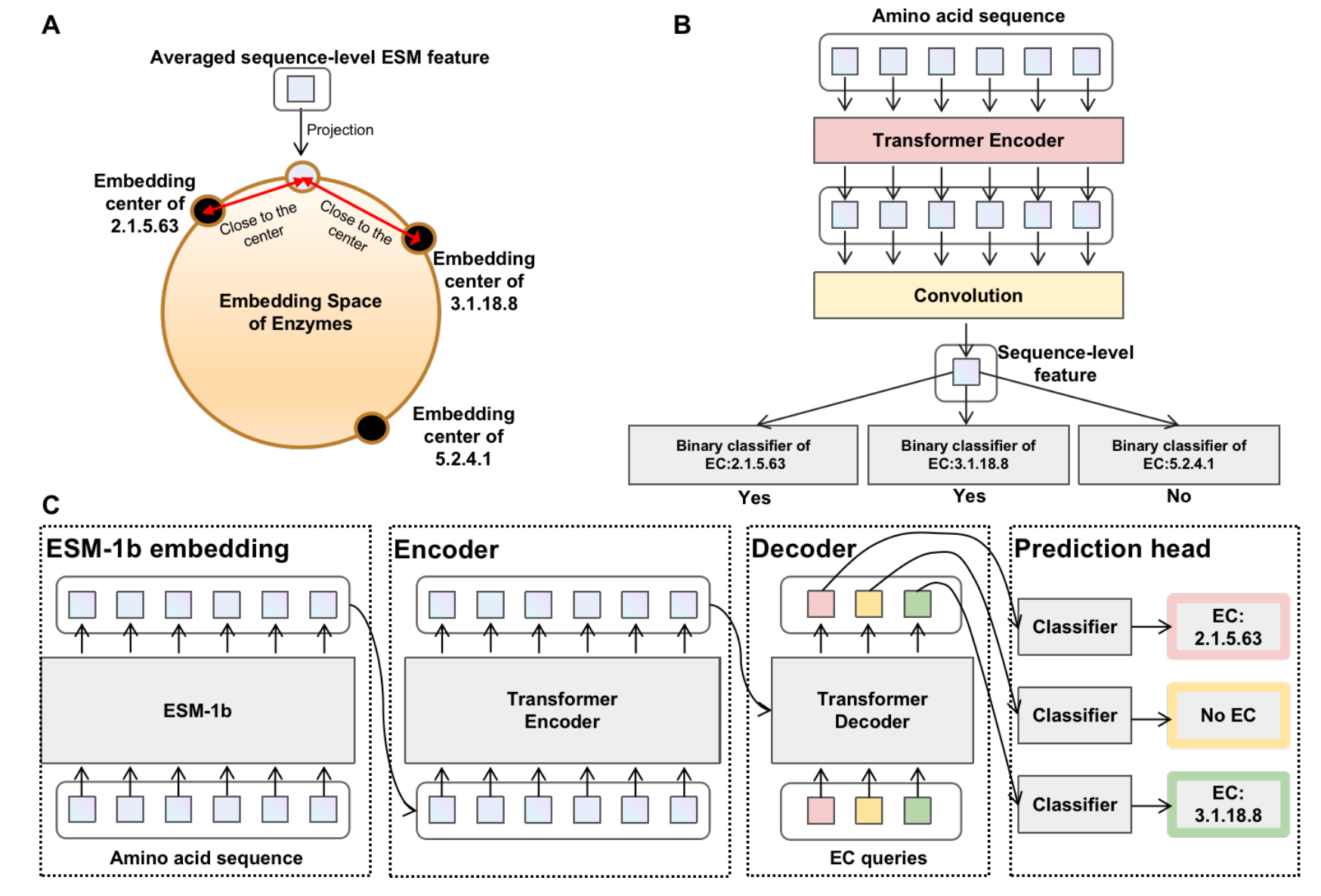
Interpretable Enzyme Function Prediction via Residue-Level Detection
Zhao Yang, Bing Su, Jiahao Chen, Ji-Rong Wen arXiv / ICLR LMRL Workshop, 2025 Paper / code We propose ProtDETR, a novel framework that reframes enzyme function prediction as an object detection problem by identifying active sites as "objects". |
|
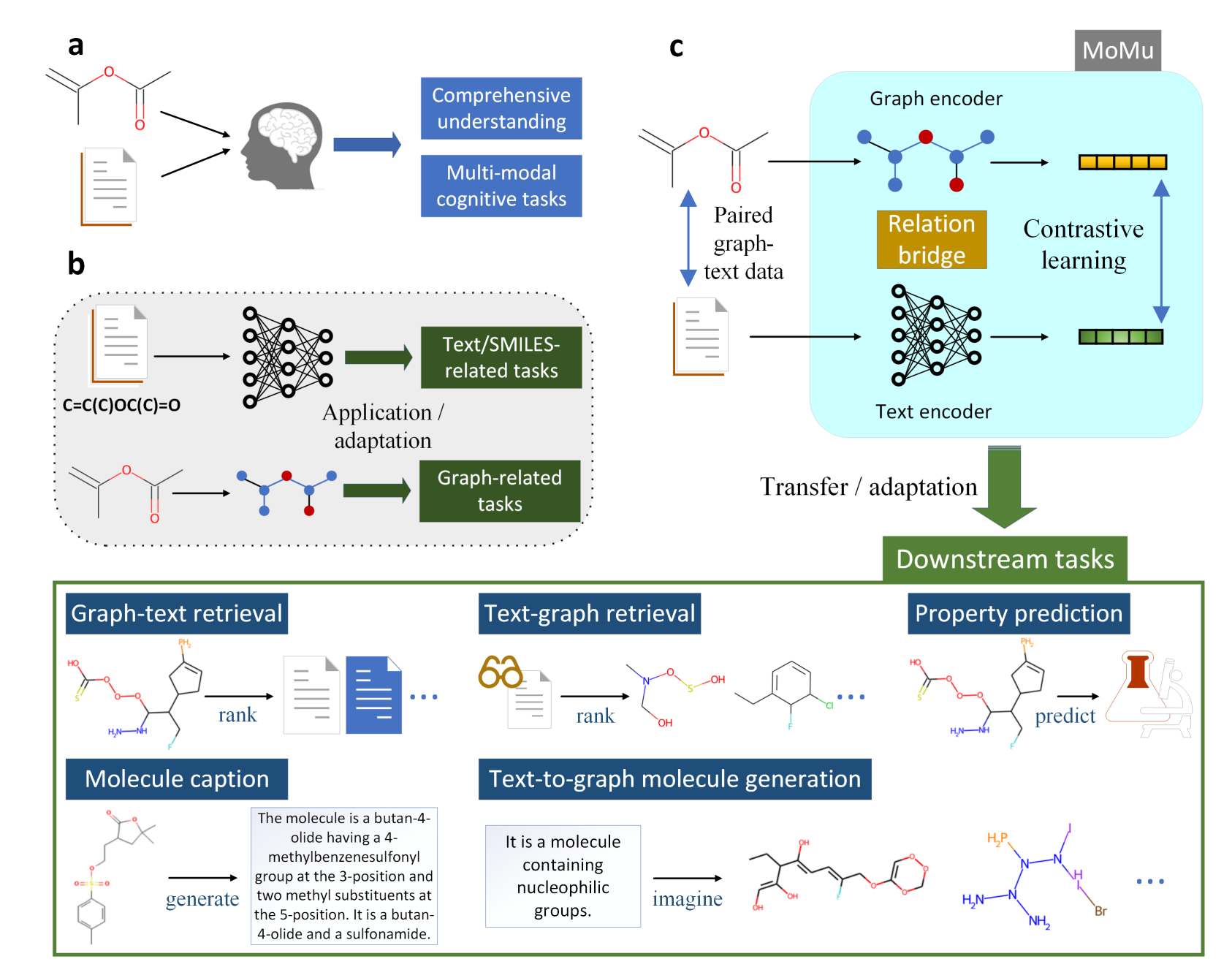
A Molecular Multimodal Foundation Model Associating Molecule Graphs with Natural Language
Bing Su, Dazhao Du, Zhao Yang, Yujie Zhou, Jiangmeng Li, Anyi Rao, Hao Sun, Zhiwu Lu, Ji-Rong Wen arXiv, 2022 Paper / code We propose MoMu, a foundation model associating molecular graphs with natural language, enabling multimodal understanding in molecular science. |
2026 |
|
|
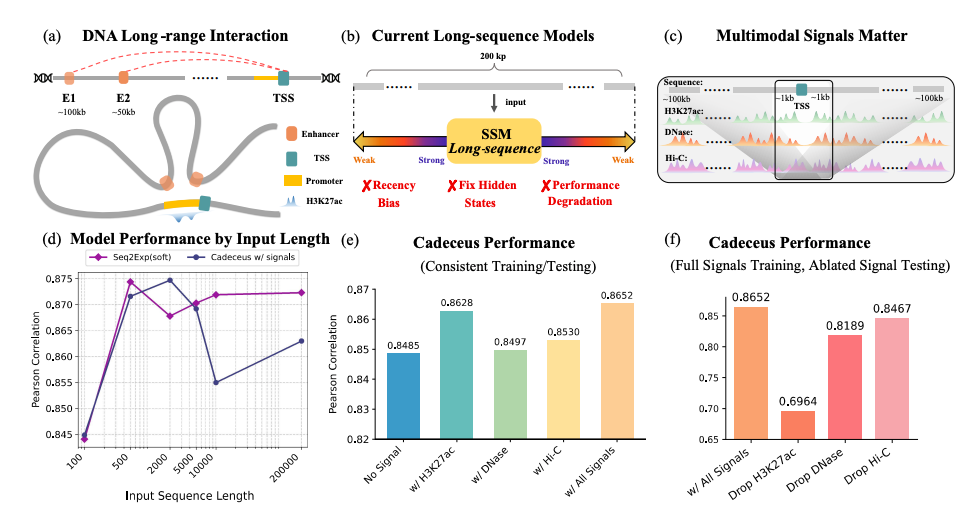
Extending Sequence Length is Not All You Need: Effective Integration of Multimodal Signals for Gene Expression Prediction
Zhao Yang*, Yi Duan*, Jiwei Zhu, Ying Ba, Chuan Cao, Bing Su ICLR, 2026 (Oral) Paper / code We propose Prism, a multimodal framework that effectively integrates diverse epigenomic signals with DNA sequences for gene expression prediction, achieving state-of-the-art performance. |
2025 |
|
|
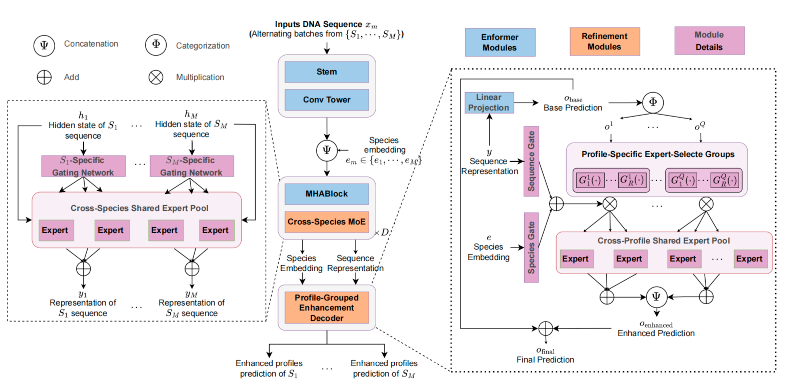
SPACE: Your Genomic Profile Predictor is a Powerful DNA Foundation Model
Zhao Yang*, Jiwei Zhu*, Bing Su ICML, 2025 Paper / code We leverage MoE to better model cross-species and cross-genomic profile information in genomic data and hence make sequence-to-function genomic models to be powerful DNA foundation models. |
|
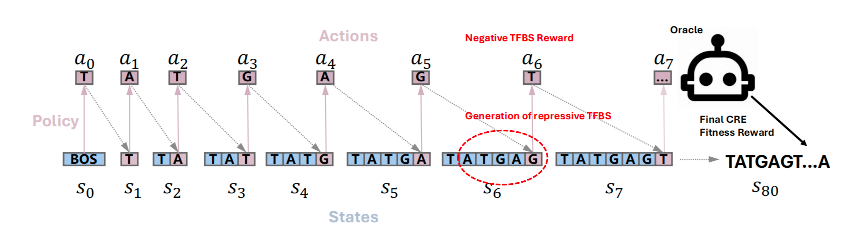
Regulatory DNA Sequence Design with Reinforcement Learning
Zhao Yang, Bing Su, Chuan Cao, Ji-Rong Wen ICLR, 2025 Paper / code We propose a reinforcement learning framework for designing regulatory DNA sequences using transcription factor binding site rewards. |
2023 |
|
|
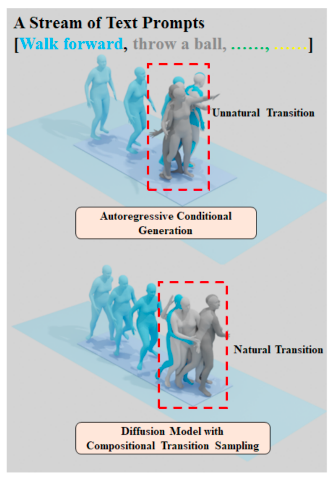
Synthesizing Long-Term Human Motions with Diffusion Models via Coherent Sampling
Zhao Yang, Bing Su, Ji-Rong Wen ACM Multimedia, 2023 Paper / code A text-driven diffusion-based framework for synthesizing long-term human motions, achieving coherent and realistic sequences. |
|
Template from Jon Barron. |